2021 has been the founding year of the Bojar lab (yay!), which means lots of building up (people, equipment, workflows, etc.). Nevertheless we were busy bees and got quite a lot to show for it! In addition to published papers (see below), we also received an NMMP Scientific Network Facilitation Grant from the National Molecular Medicine Program (NMMP), an umbrella organization connecting different Wallenberg centers in Sweden. With the publication of LectinOracle, we were also part of the “Rising Stars” series highlighting exceptionally promising early-career researchers, launched by the journal Advanced Science. Let’s see what 2022 holds!
1) Gene switch for L-glucose-induced biopharmaceutical production in mammalian cells
Published in Biotechnology and Bioengineering, this work demonstrates the control of human gene expression with a rare monosaccharide that is absent from human life: the mirror image form of glucose. Read more about it here.
2) DeepConnection: Classifying Relationship State from Images of Romantic Couples
Using a convolutional neural network, we have here analyzed the relationship satisfaction of romantically involved couples from still images and videos. This work has been published in the Journal of Computational Social Science and you can read more about it here.
3) Graph Convolutional Neural Networks to Analyze Complex Carbohydrates
This one came out in Cell Reports and describes the development of our graph convolutional neural network for the analysis of complex, nonlinear glycans. When published it was the clear new state-of-the-art for applications using glycans and introduced a new type of application, the prediction of glycan-protein interactions. Read more here.


4) Glycowork: A Python package for glycan data science and machine learning
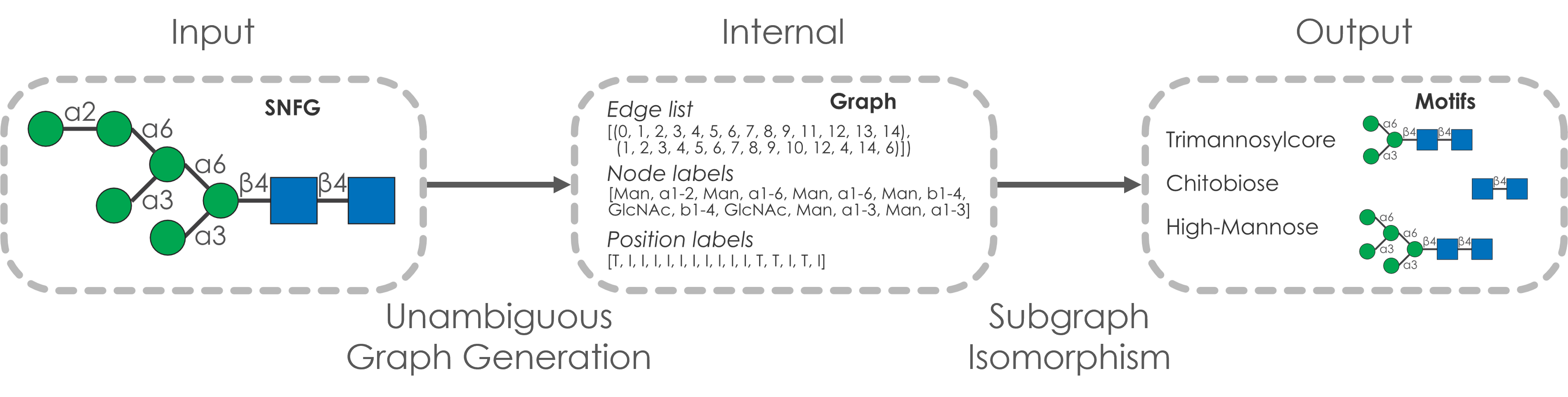
 Glycowork is our vision of democratizing glycan-focused data science and machine learning. Packed with high-level wrapper function that allow for powerful analysis routines in a single line of code as well as all the individual functions for experienced users, it is a continuously evolving project for us. Currently, it is in version 0.3, with many new features planned for 0.4. The full documentation can always be found here and I’d like to point out that glycowork also stores our continuously curated public datasets (such as ~31,500 species-specific glycans or >550,000 protein-glycan interactions). Glycowork is published in the journal Glycobiology.
Glycowork is our vision of democratizing glycan-focused data science and machine learning. Packed with high-level wrapper function that allow for powerful analysis routines in a single line of code as well as all the individual functions for experienced users, it is a continuously evolving project for us. Currently, it is in version 0.3, with many new features planned for 0.4. The full documentation can always be found here and I’d like to point out that glycowork also stores our continuously curated public datasets (such as ~31,500 species-specific glycans or >550,000 protein-glycan interactions). Glycowork is published in the journal Glycobiology.
5) Construction of caffeine-inducible gene switches in mammalian cells
Published in Methods in Molecular Biology, this book chapter details the generation of caffeine-inducible gene expression in human cells, as pioneered here.
6) The role of fucose-containing glycan motifs across taxonomic kingdoms
In this paper, we demonstrate the capabilities of glycowork by analyzing the role of a glycan feature across many different contexts: the addition of fucose or fucosylation. In Frontiers in Molecular Biosciences, we describe the taxonomy-specific sequence contexts of fucose, its role in host mimicry by pathogens, and its connection to the environment of a species.

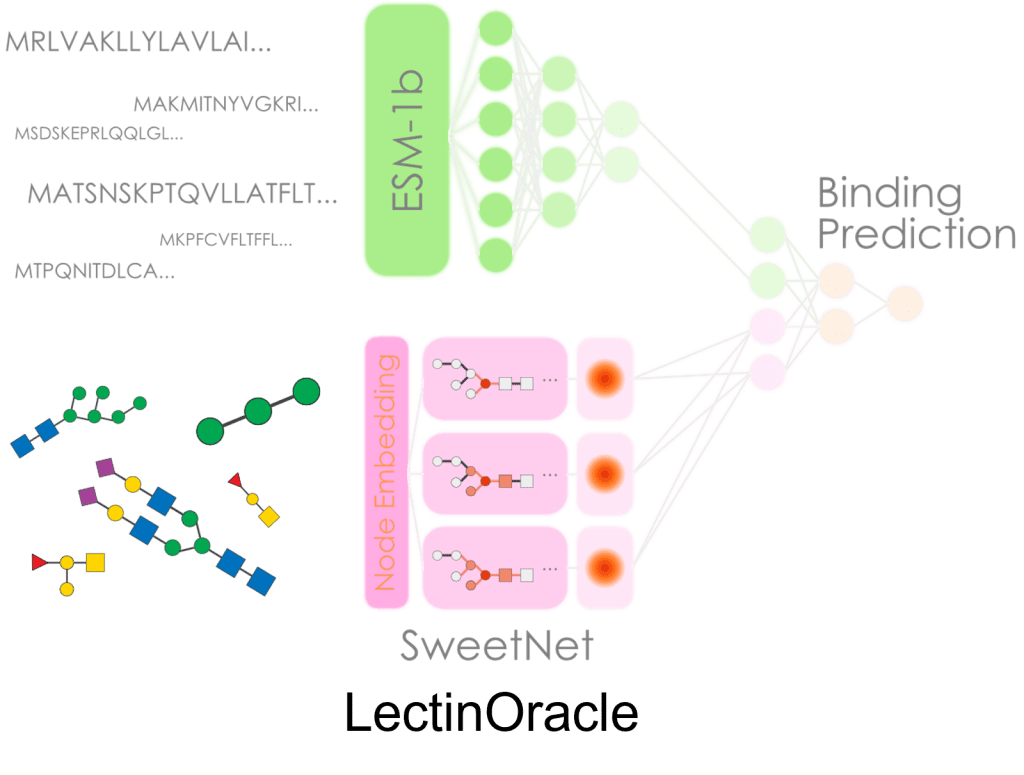
7) LectinOracle – A Generalizable Deep Learning Model for Lectin-Glycan Binding Prediction
Here, we have developed and validated a model, LectinOracle, that can very flexibly predict protein-carbohydrate binding (based on a large dataset of >550,000 protein-glycan interactions that we curated). We have for instance used this model to annotate uncharacterized proteins, study host-microbe interactions, aid directed evolution, and predict the epidemiological outcomes of viruses. Notably, LectinOracle can also be easily used within glycowork. Published in Advanced Science, LectinOracle was selected for a Frontispiece cover, highlighted in the Editorial, and part of the “Rising Stars” series highlighting exceptionally promising early-career researchers.

8) A Useful Guide to Lectin Binding: Machine-Learning Directed Annotation of 57 Unique Lectin Specificities
Here, we have used a combination of rule-based machine learning and expert curation to detail the complex binding specificities of the most commonly used glycan-binding proteins. In this process, we have uncovered many unexpected binding characteristics and present a comprehensive resource for scientists. Currently only available on bioRxiv, this work will soon be available in journal format as well…
